Research Activities
The Laboratory's main activities are:
- characterising metabolic profiles (phenotypes) of living organisms, both plants and animals;
- characterising metabolic profiles of microbial, plant, animal and human cells in relation to genetic mutations or biotic/abiotic perturbations;
- identifying new biomarkers to improve clinical diagnosis and prognosis;
- identifying new metabolic targets in various human diseases;
- assessing the effect of pharmacological, dietary or behavioural interventions on human and animal health;
- evaluating potential side effects of pharmacological treatments;
- studying interactions between diet, gut microbiota and metabolism in physiological and pathological states;
- personalising pharmacological and/or nutritional interventions based on an individual's metabolic phenotype (Precision Medicine and Precision Nutrition);
- carrying out integrated characterisation of plant-derived foods with respect to origin, pedoclimatic and agronomic conditions;
- identifying bioactive compounds in foods;
- designing functional foods and evaluating their impact on health;
- evaluating food processing stages for process optimisation;
- evaluating phases of processes involved in the agricultural circular economy.
The team managing NMLab is available to discuss potential experiments and to establish collaborations that may include advanced data analysis and interpretation.
Metabolomics
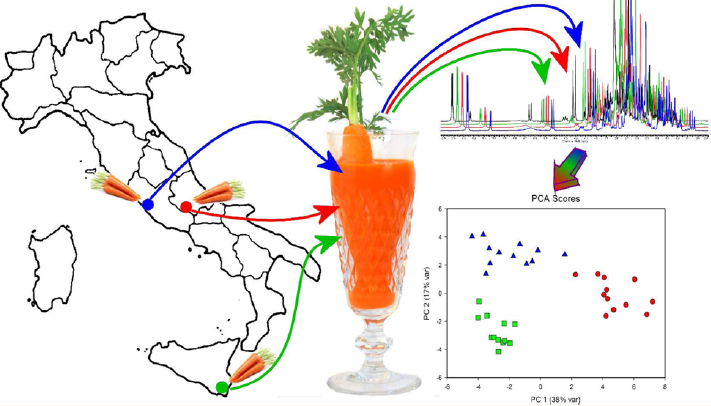
Metabolomics is the characterisation of the complete set of metabolites in a matrix of plant or animal origin, highlighting its variation according to the physiological, developmental or pathological state of a cell, tissue, organ or organism. This characterisation makes it possible to reconstruct the network of metabolic processes in the system under study.
Metabolomics has applications in many areas, including human and animal health, biomarker discovery, drug discovery and development, plant biology, microbiology, food chemistry and environmental monitoring. It enables a deeper understanding of biological systems and a more detailed description of biochemical processes and their interconnections.
It is important to emphasise that, at present, NMR-based metabolomics performed at NMLab cannot be considered a diagnostic tool or a clinical test. It is instead a scientific investigation on populations of subjects to identify metabolic traits that may distinguish different groups under specific conditions. It is therefore not a diagnostic exam and cannot be applied to single cases.
NMR spectroscopy is the preferred technique for structural characterisation and quantification of molecules in complex mixtures, which can be performed in a single spectrum without pre-treatment. It offers major advantages over other analytical techniques, including ease of quantification, direct identification of metabolites and the ability to detect unexpected metabolites. NMR analysis of a sample enables the simultaneous detection of low molecular weight metabolites present in a solution at concentrations above 10-6 M.
NMR-based metabolomics analysis mainly relies on multivariate statistical methods applied to datasets obtained from NMR spectra.
NMR Experiments
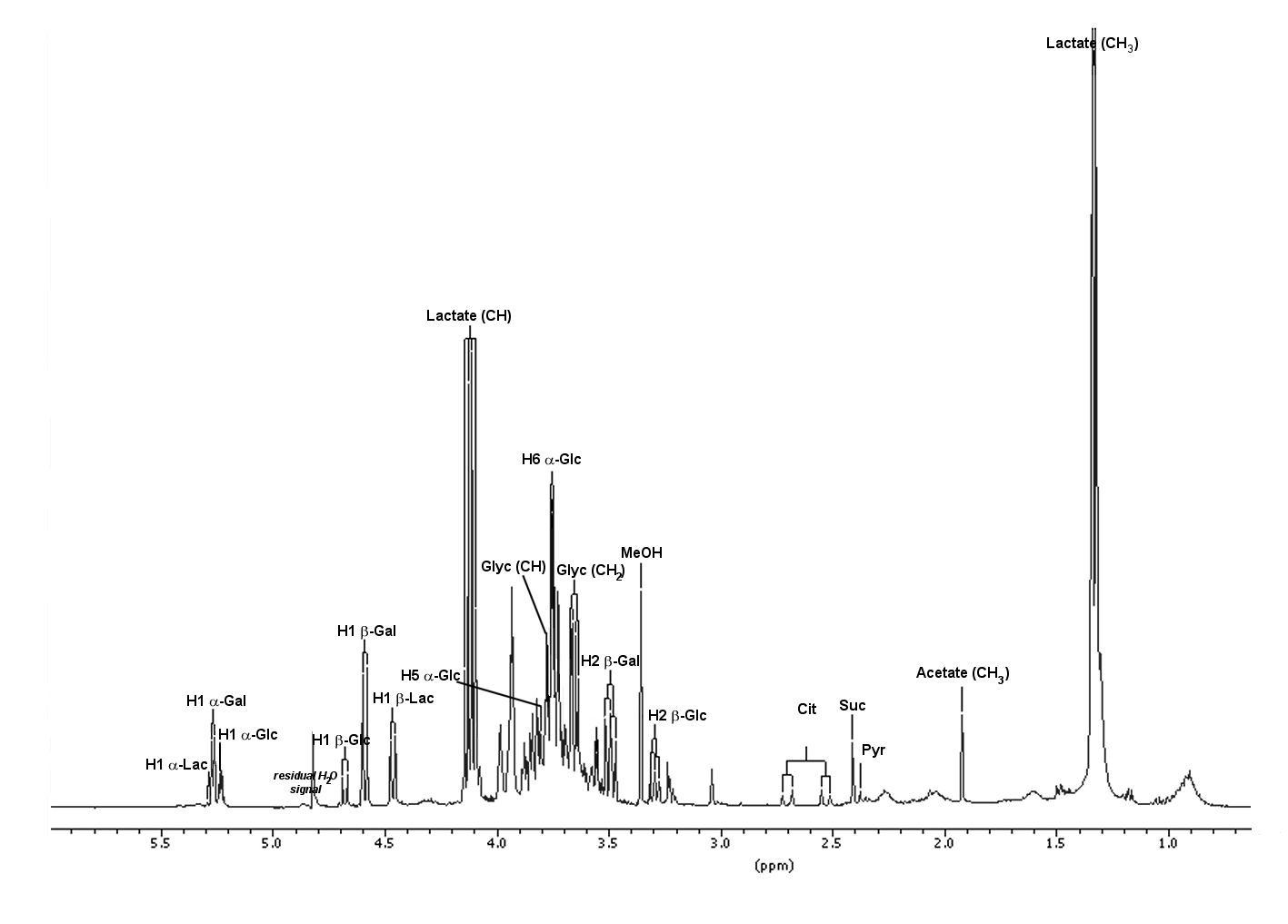
NMR spectra consist of a series of resonance signals, each originating from magnetically equivalent nuclei (hydrogen nuclei in 1H NMR spectroscopy). Their position in the spectrum (chemical shift) is characteristic of the chemical group the nucleus belongs to, as well as its chemical environment, while their intensities (areas) are proportional to the concentration of the chemical group and thus of the molecule.
Since every metabolite contains more than one hydrogen-bearing chemical group, each metabolite is represented by multiple NMR signals. A 1D 1H spectrum therefore produces a crowded set of resonance signals, an instantaneous snapshot of all metabolites in the mixture, often described as its fingerprint or metabolic fingerprint. Signal assignment can be greatly simplified using 2D NMR experiments, which expand the complexity of the 1D spectrum into a second dimension.
Data Analysis
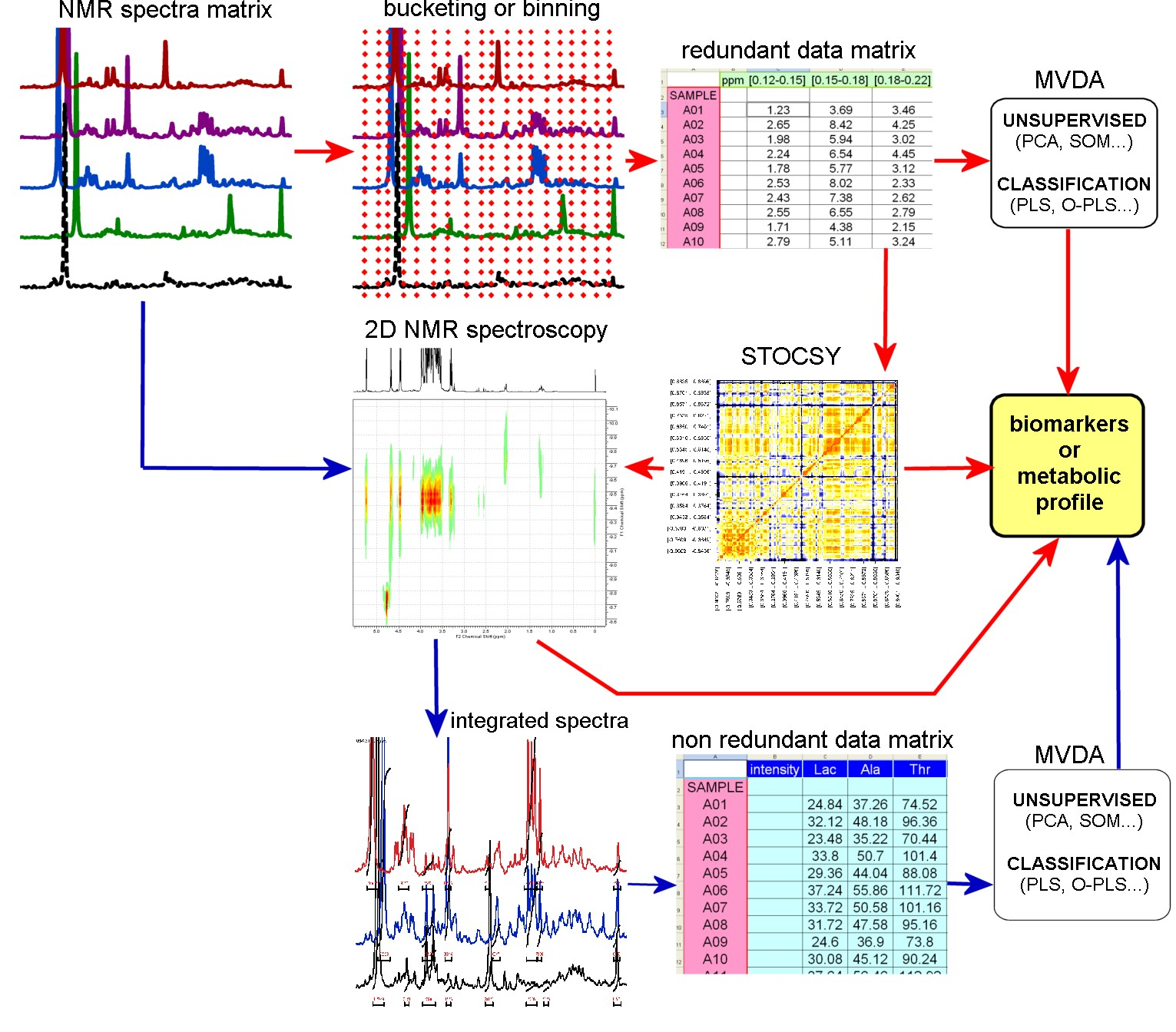
Quantification is the step that transforms a set of NMR spectra into a data matrix. This objective can be achieved through two complementary approaches: assigning each (or most) signals to the correct metabolite and measuring peak intensities (targeted analysis), or processing spectra as is regardless of prior assumptions, relying on correlation structures embedded in the intrinsic redundancy of the 1D 1H spectrum (pattern recognition).
Both targeted analysis and pattern recognition require assigning resonance signals to their metabolite sources. This procedure relies on operator expertise and may use a variety of 2D NMR techniques, reference spectra and literature data. Both contribute to generating metabolic profiles and detecting biomarkers that characterise the biological processes occurring in the system.
Once these steps are completed, multivariate techniques are applied to detect - in an unsupervised manner - intrinsic correlations and/or outliers in the dataset (commonly via Principal Component Analysis, PCA) or - in a supervised manner - to use known external variables (e.g., Partial Least Squares, PLS; Orthogonal-PLS; Multiple Correspondence Analysis, MCA) or for classification (PLS, SIMCA, Multiway analysis, etc.).
